Families & Caregivers
Hope for a Cure
Angelman syndrome (AS) is a monogenic disorder, meaning it results from the dysfunction of a single gene. The gene responsible for AS, UBE3A, is unique in that it is imprinted, which means that the expression of the gene depends upon whether the gene was inherited through the egg (maternal) or the sperm (paternal).
In all individuals, the paternal copy of the UBE3A gene is turned off or silenced in most brain cells. This means that the maternal copy of the UBE3A gene is responsible for making the UBE3A protein in the brain. However, when the maternal copy of UBE3A is missing or not functional, UBE3A protein is no longer produced in the brain. This absence of functional UBE3A protein affects typical brain function and causes Angelman Syndrome.

Understanding Angelman Syndrome and Its Treatment Possibilities
In AS, the unique nature of the UBE3A gene offers a promising opportunity for potential therapies. Due to UBE3A's imprinting, individuals living with AS usually have a functional paternal copy of the UBE3A gene. This presents an exciting possibility: the paternal UBE3A gene could potentially be "turned on" to enable the production of UBE3A protein. As a result, there are therapeutic approaches that could address the underlying cause of AS that may not be possible in other similar genetic conditions. Traditional strategies used in genetic disorders, such as gene replacement therapy or enzyme replacement therapy, remain potential therapeutic options as well.
At FAST, we think about therapeutic development by classifying into pillars.
Pillar 1: Replace Mom’s UBE3A
Pillar 2: Turn on Dad’s UBE3A
Pillar 3: Explore Adjacent Genes
Pillar 4: Target Downstream Pathways
Pillar 5: Accelerating and Preparing for Clinical Trials
Are There Any Treatments for Angelman Syndrome?
Currently, treatments for AS are generally targeted to the symptoms each individual is experiencing. For example, individuals experiencing seizures should receive treatment(s) directed at reducing or eliminating the seizures. In 2022, a consensus statement on care in AS was published, offering valuable guidance to health care providers about managing symptoms and preventative care for individuals living with Angelman syndrome.
There is currently no cure for Angelman syndrome. However, research is actively progressing, aimed at developing potential therapeutics to improve symptoms and addressing the underlying cause of AS, the missing UBE3A protein in the brain. The Drug Development Pipeline provides information on potential therapeutics that are still in early stage development as well as current ongoing clinical trials.
How does FAST support the development of transformative therapies?
FAST is dedicated to funding innovative research and high-risk, high-reward research aimed at accelerating the development of new therapeutic strategies. By investing in cutting-edge science, FAST has played a pivotal role in advancing the field to the point where potential meaningful treatments for Angelman syndrome are closer than ever before.
Moreover, the work being carried out today has the potential to significantly shape the future of genomic medicine, offering hope for improved treatments and outcomes. These efforts not only aim to improve the quality of life for individuals living with Angelman syndrome but also have the potential to positively impact the lives of thousands of individuals worldwide who are affected by similar neurogenetic conditions.
FAST and other partner organizations are also working at the federal and state level to advocate for policies that have the potential to positively impact the lives of families living with AS, including increasing the federal focus and research investment in AS and supporting FDA flexibility and appropriate speed in review of potential therapies. Learn more about AS advocacy Hope in Action.
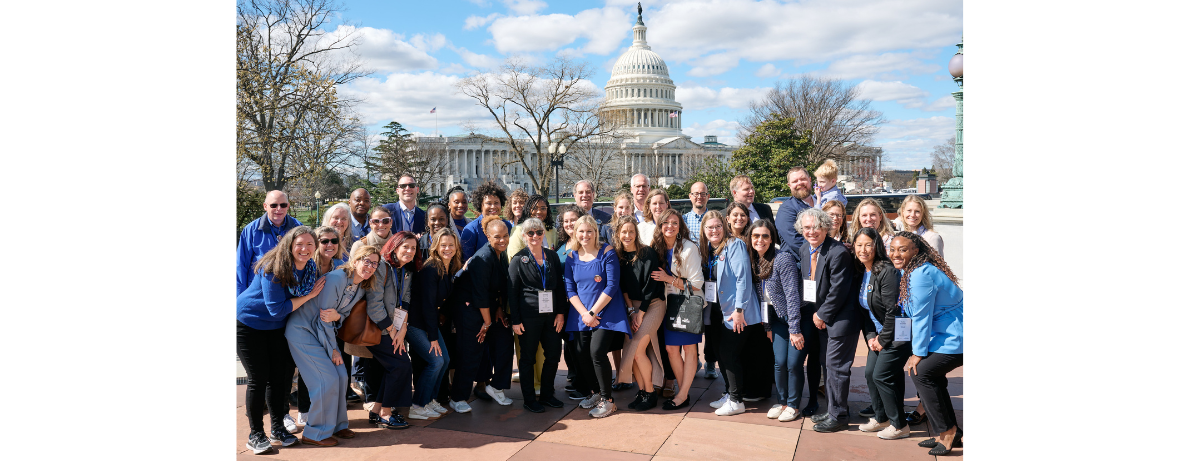

What’s Next?
The field of Angelman syndrome research is moving rapidly. Families, caregivers, and advocates can get involved and stay informed about the latest developments by:
Connecting with FAST and other advocacy organizations to stay up-to-date on breakthroughs.
Checking out the Drug Development Pipeline for an overview of potential therapeutics and where they are in development.
Learning more about advocacy for AS at the Hope in Action Center: Progress Through Advocacy site
Joining our Cure Angelman Network
We truly believe that a cure for Angelman syndrome is possible. To learn more about advancements in AS research, visit our Drug Development Pipeline.